The current research in the lab can be loosely grouped into three areas: (1) Studying DNA-protein interactions using smFRET (in-vitro) and live-cell imaging, (2) super-resolution food microscopy, and (3) methods development for single-molecule microscopy.
1. DNA-protein interactions (in vitro and in vivo)
The genetic information of any organism is stored in a very remarkable molecule known as DNA. Since DNA is such an essential part of every living cell, many different enzymes interact either directly or indirectly with it.
The first project is on plant related proteins belonging to the AUXIN RESPONSE FACTOR (ARF) family that represent DNA-binding transcription factors. Here we aim to develop and utilise new fluorescence-based assays to quantify the binding kinetics of various ARF-DBD proteins to DNA whilst monitoring structural changes of the DNA and the associated proteins.
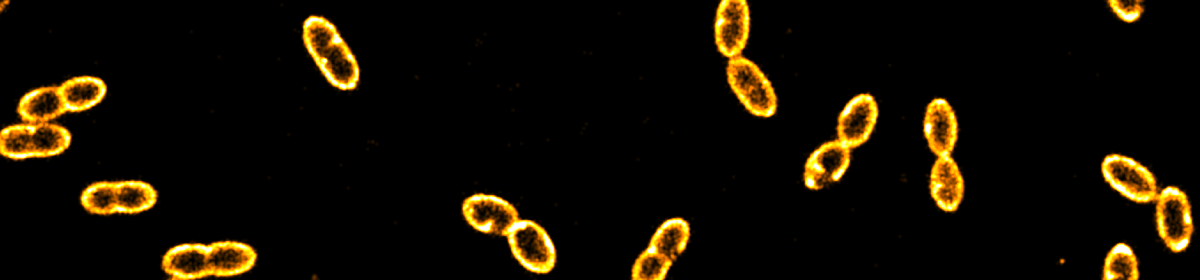
The second project focuses on the visualisation of CRISPR-Cas in live bacteria. Starting with a food-related strain of Gram-positive Lactococcus lactis, we are currently working on monitoring target search of various CRISPR variants in Gram-negative Escherichia coli.

2. Super-resolution food microscopy
In order to feed the world population in a more sustainable manner, many of the current food products will have to be reformulated, ideally by using plant-derived materials. Together with industrial stake holders we are working on illuminating the spatial and temporal organisation and involved in the oxidation of food products such as mayonnaise and yoghurt.

3. Development of single-molecule techniques
We have been working on specially designed nanofluidic devices that combine the advantages of diffusion-based, confocal microscopy (providing high time resolution for studying fast conformational changes) and camera-based microscopy (with the ability to monitor many individual molecules in parallel). We further develop and apply fluidic devices for the continuous monitoring of bacterial cells.
A second direction is the development of new tools for spectrally resolved super-resolution microscopy.