Fluorescence microscopy is an extremely powerful and versatile technique contributing to many areas of the life sciences. To increase the affordability and accessibility of single-molecule microscopy, our group frequently releases information on both hardware and software. We frequently post new software on Github here.
(Hardware) The miCube open microscopy framework
The miCube is an open and modular hardware framework aiming for cost effectiveness, modularity, customisability, openness, stability and throughput. Publication here, project on Github here.
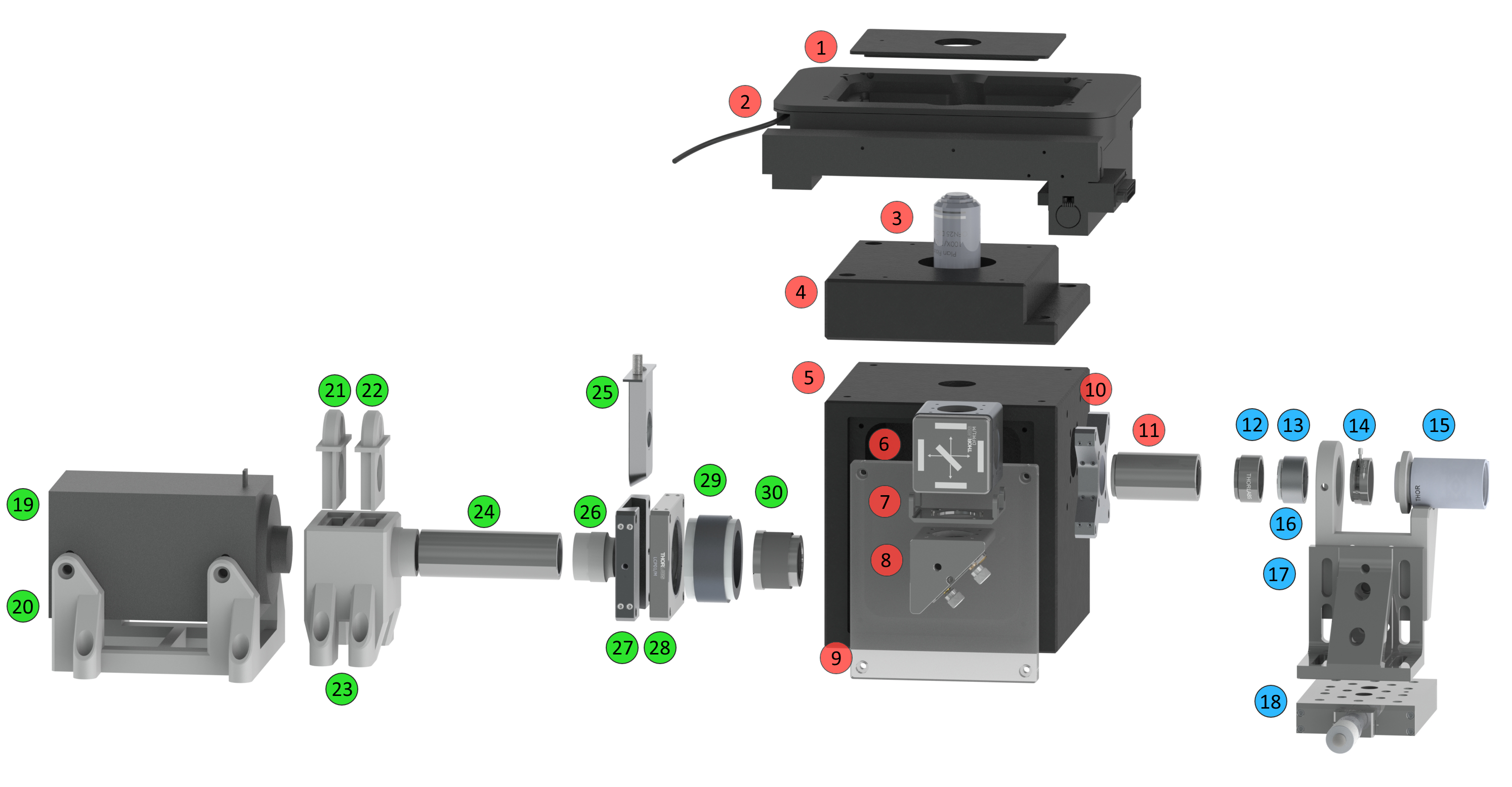
(Hardware/software) SMILE: single-molecule imaging laser engine
The concept of the SMILE project is to develop a low cost laser controller based on an Arduino NANO which is capable to operate four low cost, in this case JLaserTrack LDM series, lasers/drivers. All necessary information (software, hardware list, etc) can be found here.
(Software) pSMLM: phasor-based single-molecule localisation microscopy
Phasor-based single-molecule localisation microscopy (pSMLM-3D) is a fast and model-free 2D and 3D single-molecule localization algorithm that allows millions of localizations per second to be calculated on CPUs with localization accuracies in line with the most accurate algorithms currently available. Read more about pSMLM-3D here, or start using it directly in ImageJ/Fiji by getting the pSMLM-3D ThunderSTORM plugin. We recently extended pSMLM to include the analysis of engineered point spread functions. Publication here, software (Matlab) here.
(Software) FTM2: fast temporal median filtering
Temporal median filtering is a simple technique to estimate and correct background intensity in noisy camera recordings. Compared to previous iterations, our version enabled extra features like more bit-depth support and larger-than-ram datasets, as well as being slightly faster than previous iterations. The latest version of the plugin for ImageJ/Fiji can be found here.
(Software) anaDDA: analytical diffusion distribution analysis
Single-particle tracking is an important technique in the life sciences to understand the kinetics of biomolecules. To analyse the diffusivity of, for example, fluorescently labelled proteins in live bacteria we recently developed a analytical framework that excels in analysing short tracks that consist only of a few localisations. Publication here, software (Matlab) here.