J. Hohlbein, Food Structure, 30, 100236, 2021, [link]
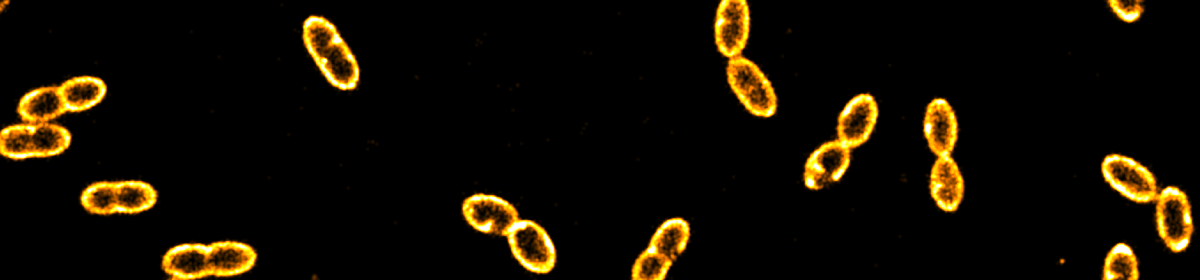
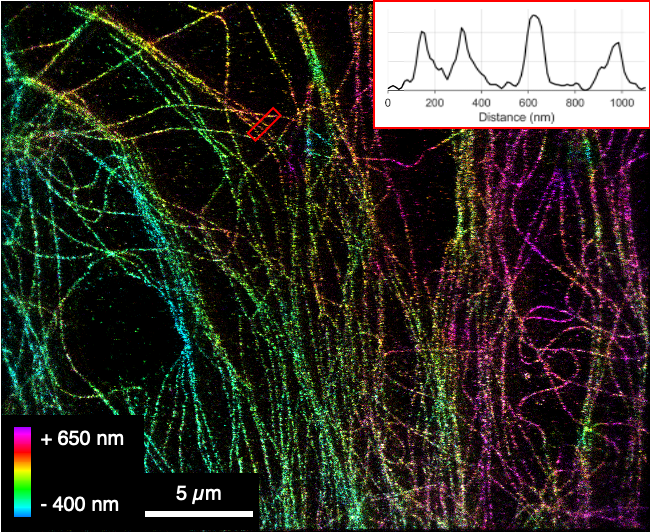
Optical microscopy is an indispensable tool to characterize the microstructure of foods at ambient conditions. Depending on both the wavelength of light used to illuminate the sample and the opening angle of the microscope objective, the achievable resolution is limited to around 200 nm. This so-called classical diffraction limit implies that smaller structural features cannot be resolved or separated from each other. As many food structures are ultimately defined by the molecular interactions of single proteins or single molecules, the classical resolution is insufficient to reveal structural details in the (tens of) nanometer range. Intriguingly, recent advancements in imaging techniques originating mostly in the (biomedical) life sciences have been closing the gap, pushing the resolution towards true molecular resolution. In this perspective, we want to highlight some of these emerging techniques and provide an outlook on potential future applications.