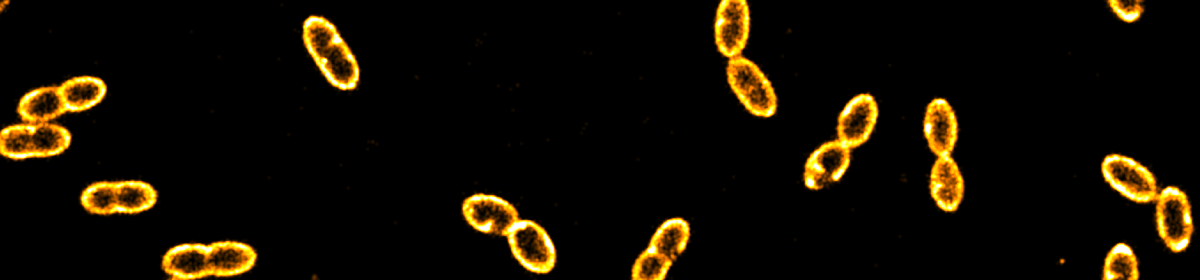
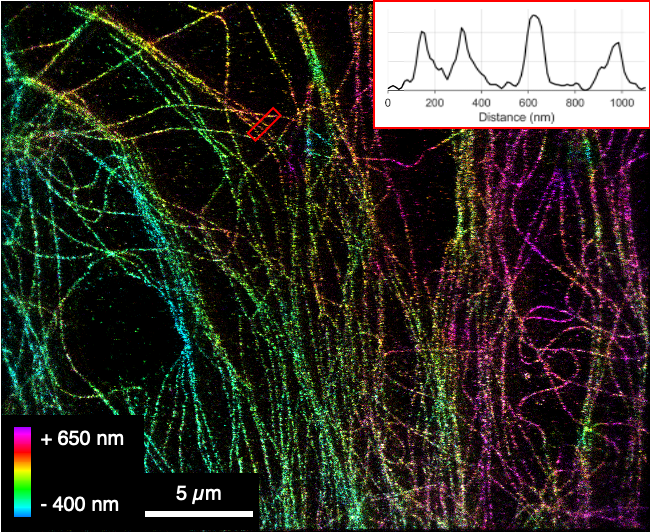
Turbidity poses a major challenge for the microscopic characterization of many food systems. In these systems, local mismatches in refractive indices can cause reflection, absorption and scattering of incoming as well as outgoing light leading to significant image deterioration along sample depth. To mitigate the issue of turbidity and to increase the achievable optical resolution, we combined adaptive optics (AO) with single-molecule localization microscopy (SMLM). Building on our previously published open hardware microscopy framework, the miCube, we first added a deformable mirror to the detection path. This element enables both the compensation of aberrations directly from single-molecule data and, by further modulating the emission wavefront, the introduction of various point spread functions (PSFs) to enable SMLM in three dimensions. We further added a top hat beam shaper to the excitation path to obtain an even illumination profile across the field of view (FOV). As a model system for a non-transparent food colloid in which imaging in depth is challenging, we designed an oil-in-water emulsion in which phosvitin, a ferric ion binding protein present in from egg yolk, resides at the oil water interface. We targeted phosvitin with fluorescently labelled primary antibodies and used PSF engineering to obtain 2D and 3D images of phosvitin covered oil droplets with sub 100 nm resolution. Droplets with radii as low as 200 nm can be discerned, which is beyond the range of conventional confocal light microscopy. Our data indicated that in the model emulsion phosvitin is homogeneously distributed at the oil-water interface. With the possibility to obtain super-resolved images in depth of nontransparent colloids, our work paves the way for localizing biomacromolecules at colloidal interfaces in heterogeneous food emulsions.
Category: open access
Published: Single-molecule localization microscopy as an emerging tool to probe multiscale food structures
J. Hohlbein, Food Structure, 30, 100236, 2021, [link]
Optical microscopy is an indispensable tool to characterize the microstructure of foods at ambient conditions. Depending on both the wavelength of light used to illuminate the sample and the opening angle of the microscope objective, the achievable resolution is limited to around 200 nm. This so-called classical diffraction limit implies that smaller structural features cannot be resolved or separated from each other. As many food structures are ultimately defined by the molecular interactions of single proteins or single molecules, the classical resolution is insufficient to reveal structural details in the (tens of) nanometer range. Intriguingly, recent advancements in imaging techniques originating mostly in the (biomedical) life sciences have been closing the gap, pushing the resolution towards true molecular resolution. In this perspective, we want to highlight some of these emerging techniques and provide an outlook on potential future applications.
Published: Integrating engineered point spread functions into the phasor-based single-molecule localization microscopy framework
K.J.A. Martens, A. Jabermoradi, S. Yang, and J. Hohlbein, Methods, 193, 2021, [link], previously on bioRxiv [link]
In single-molecule localization microscopy (SMLM), the use of engineered point spread functions (PSFs) provides access to three-dimensional localization information. The conventional approach of fitting PSFs with a single 2-dimensional Gaussian profile, however, often falls short in analyzing complex PSFs created by placing phase masks, deformable mirrors or spatial light modulators in the optical detection pathway. Here, we describe the integration of PSF modalities known as double-helix, saddle-point or tetra-pod into the phasor-based SMLM (pSMLM) framework enabling fast CPU based localization of single-molecule emitters with sub- pixel accuracy in three dimensions. For the double-helix PSF, pSMLM identifies the two individual lobes and uses their relative rotation for obtaining z-resolved localizations. For the analysis of saddle-point or tetra-pod PSFs, we present a novel phasor-based deconvolution approach entitled circular-tangent pSMLM. Saddle-point PSFs were experimentally realized by placing a deformable mirror in the Fourier plane and modulating the incoming wavefront with specific Zernike modes. Our pSMLM software package delivers similar precision and recall rates to the best-in-class software package (SMAP) at signal-to-noise ratios typical for organic fluorophores and achieves localization rates of up to 15 kHz (double-helix) and 250 kHz (saddle-point/tetra-pod) on a standard CPU. We further integrated pSMLM into an existing software package (SMALL-LABS) suitable for single-particle imaging and tracking in environments with obscuring backgrounds. Taken together, we provide a powerful hardware and software environment for advanced single-molecule studies.
Published: Probing DNA – transcription factor interactions using single-molecule fluorescence detection in nanofluidic devices
M. Fontana, Š. Ivanovaite , S. Lindhoud, E. van der Wijk, K. Mathwig, W. van den Berg, D. Weijers, and J. Hohlbein, Advanced Biology, 2100953, 2021, [link], preprint: bioRxiv, 2021, [link]
Single-molecule fluorescence detection offers powerful ways to study biomolecules and their complex interactions. Here, nanofluidic devices and camera-based, single-molecule Förster resonance energy transfer (smFRET) detection are combined to study the interactions between plant transcription factors of the auxin response factor (ARF) family and DNA oligonucleotides that contain target DNA response elements. In particular, it is shown that the binding of the unlabeled ARF DNA binding domain (ARF-DBD) to donor and acceptor labeled DNA oligonucleotides can be detected by changes in the FRET efficiency and changes in the diffusion coefficient of the DNA. In addition, this data on fluorescently labeled ARF-DBDs suggest that, at nanomolar concentrations, ARF-DBDs are exclusively present as monomers. In general, the fluidic framework of freely diffusing molecules minimizes potential surface-induced artifacts, enables high-throughput measurements, and proved to be instrumental in shedding more light on the interactions between ARF-DBDs monomers and between ARF-DBDs and their DNA response element
Published: Spatiotemporal heterogeneity of κ‐carrageenan gels investigated via single-particle-tracking fluorescence microscopy
Hydrogels made of the polysaccharide κ-carrageenan are widely used in the food and personal care industry as thickeners or gelling agents. These hydrogels feature dense regions embedded in a coarser bulk network, but the characteristic size and behavior of these regions has remained elusive. Here, we use single-particle-tracking fluorescence microscopy (sptFM) to quantitatively describe κ-carrageenan gels. Infusing fluorescent probes into fully gelated κ-carrageenan hydrogels resulted in two distinct diffusional behaviors. Obstructed self-diffusion of the probes revealed that the coarse network consists of κ-carrageenan strands with a typical diameter of 3.2 ± 0.3 nm leading to a nanoprobe diffusion coefficient of ~1-5∙10^-12 m2/s. In the dense network regions, we found a fraction with a largely decreased diffusion coefficient of ~1∙10^-13 m2/s. We also observed dynamic exchange between these states. The computation of spatial mobility maps from diffusional data indicated that the dense network regions have a characteristic diameter of ~1 µm and are itself mobile on the seconds-to-minutes timescale. sptFM provides an unprecedented view on spatiotemporal heterogeneity of hydrogel networks, which we believe bears general relevance for understanding transport and release of both low- and high molecular weight solutes.

Pre-print: Analyzing engineered point spread functions using phasor-based single-molecule localization microscopy
The point spread function (PSF) of single molecule emitters can be engineered in the Fourier plane to encode three-dimensional localization information, creating double-helix, saddle-point or tetra-pod PSFs. Here, we describe and assess adaptations of the phasor-based single-molecule localization microscopy (pSMLM) algorithm to localize single molecules using these PSFs with sub-pixel accuracy. For double-helix, pSMLM identifies the two individual lobes and uses their relative rotation for obtaining z-resolved localizations, while for saddle-point or tetra-pod, a novel phasor-based deconvolution approach is used. The pSMLM software package delivers similar precision and recall rates to the best-in-class software package (SMAP) at signal-to-noise ratios typical for organic fluorophores. pSMLM substantially improves the localization rate by a factor of 2 – 4x on a standard CPU, with 1-1.5·104 (double-helix) or 2.5·105 (saddle-point/tetra-pod) localizations/second.

Published: Substrate conformational dynamics facilitate structure-specific recognition of gapped DNA by DNA polymerase
, Nucleic Acid Research, 47, 10788, 2019, [link]
DNA-binding proteins utilise different recognition mechanisms to locate their DNA targets; some proteins recognise specific DNA sequences, while others interact with specific DNA structures. While sequence-specific DNA binding has been studied extensively, structure-specific recognition mechanisms remain unclear. Here, we study structure-specific DNA recognition by examining the structure and dynamics of DNA polymerase I Klenow Fragment (Pol) substrates both alone and in DNA–Pol complexes. Using a docking approach based on a network of 73 distances collected using single-molecule FRET, we determined a novel solution structure of the single-nucleotide-gapped DNA–Pol binary complex. The structure resembled existing crystal structures with regards to the downstream primer-template DNA substrate, and revealed a previously unobserved sharp bend (∼120°) in the DNA substrate; this pronounced bend was present in living cells. MD simulations and single-molecule assays also revealed that 4–5 nt of downstream gap-proximal DNA are unwound in the binary complex. Further, experiments and coarse-grained modelling showed the substrate alone frequently adopts bent conformations with 1–2 nt fraying around the gap, suggesting a mechanism wherein Pol recognises a pre-bent, partially-melted conformation of gapped DNA. We propose a general mechanism for substrate recognition by structure-specific enzymes driven by protein sensing of the conformational dynamics of their DNA substrates.

Publication: Visualisation of dCas9 target search in vivo using an open-microscopy framework
K.J.A. Martens, S. van Beljouw, S. van der Els, J.N.A. Vink, S. Baas, G.A. Vogelaar, S.J.J. Brouns, P. van Baarlen, M. Kleerebezem, J. Hohlbein, Nature Communications, 10, 3552, 2019, [link]
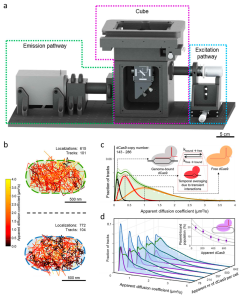
CRISPR-Cas9 is widely used in genomic editing, but the kinetics of target search and its relation to the cellular concentration of Cas9 have remained elusive. Effective target search requires constant screening of the protospacer adjacent motif (PAM) and a 30 ms upper limit for screening was recently found. To further quantify the rapid switching between DNA-bound and freely-diffusing states of dCas9, we developed an open-microscopy framework, the miCube, and introduce Monte-Carlo diffusion distribution analysis (MC-DDA). Our analysis reveals that dCas9 is screening PAMs 40% of the time in Gram-positive Lactoccous lactis, averaging 17 ± 4 ms per binding event. Using heterogeneous dCas9 expression, we determine the number of cellular target-containing plasmids and derive the copy number dependent Cas9 cleavage. Furthermore, we show that dCas9 is not irreversibly bound to target sites but can still interfere with plasmid replication. Taken together, our quantitative data facilitates further optimization of the CRISPR-Cas toolbox.
Pre-print: Direct visualization of native CRISPR target search in live bacteria reveals Cascade DNA surveillance mechanism
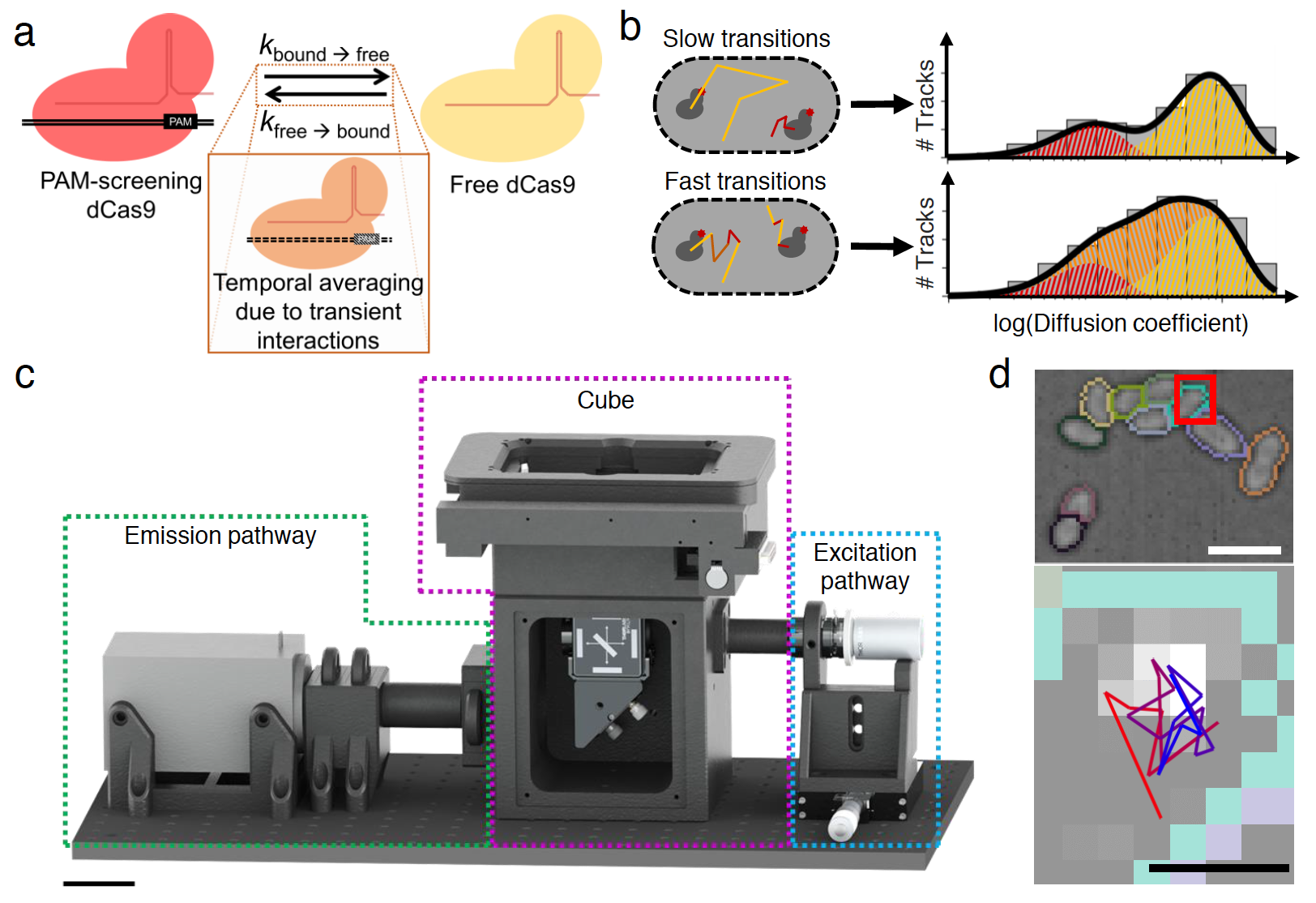
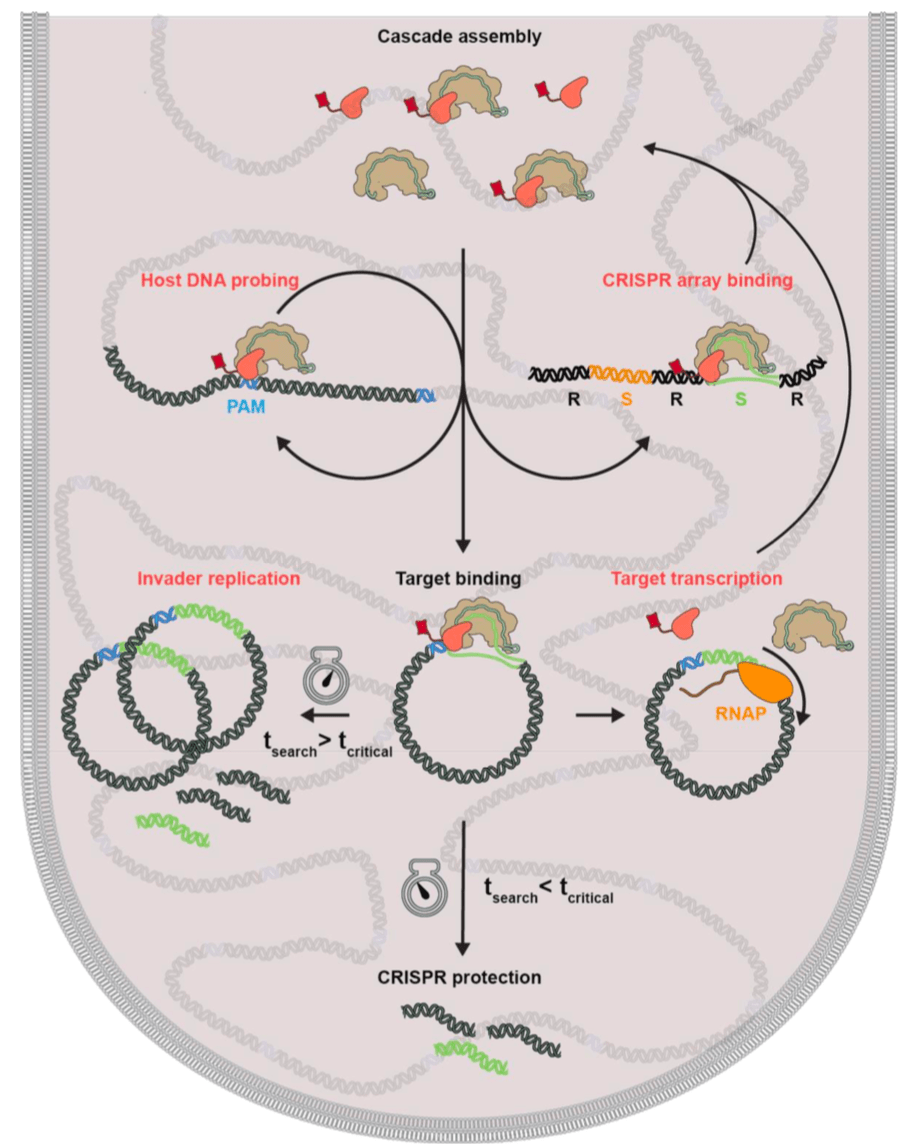
CRISPR-Cas systems encode RNA-guided surveillance complexes to find and cleave invading DNA elements. While it is thought that invaders are neutralized minutes after cell entry, the mechanism and kinetics of target search and its impact on CRISPR protection levels have remained unknown. Here we visualized individual Cascade complexes in a native type I CRISPR-Cas system. We uncovered an exponential relationship between Cascade copy number and CRISPR interference levels, pointing to a time-driven arms race between invader replication and target search, in which 20 Cascade complexes provide 50% protection. Driven by PAM-interacting subunit Cas8e, Cascade spends half its search time rapidly probing DNA (∼30 ms) in the nucleoid. We further demonstrate that target DNA transcription and CRISPR arrays affect the integrity of Cascade and impact CRISPR interference. Our work establishes the mechanism of cellular DNA surveillance by Cascade that allows the timely detection of invading DNA in a crowded, DNA-packed environment.
Pre-print: An open microscopy framework suited for tracking dCas9 in live bacteria
K.J.A. Martens, S. van Beljouw, S. van der Els, S. Baas, J.N.A. Vink, S.J.J. Brouns, P. van Baarlen, M. Kleerebezem, J. Hohlbein, bioRxiv, 2018 [link]
Super-resolution microscopy is frequently employedin the life sciences, but the number of freely accessible and affordable microscopy frameworks, especially for single particle tracking photo-activation localization microscopy (sptPALM), remains limited. To this end, we designed the miCube: a versatile super-resolution capable fluorescence microscope, which combines high spatiotemporal resolution, good adaptability, low price, and easy installation. By providing all details, we hope to enable interested researchers to build an identical or derivative instrument. The capabilities of the miCube are assessed with a novel sptPALM assay relying on the heterogeneous expression of target genes. Here, we elucidate mechanistic details of catalytically inactive Cas9 (dead Cas9) in live Lactococcus lactis. We show that, lacking specific DNA target sites, the binding and unbinding of dCas9 to DNA can be described using simplified rate constants of kbound_free = 30 – 80 s-1 and kfree_bound = 15 – 40 s-1. Moreover, after providing specific DNA target sites via DNA plasmids, the plasmid-bound dCas9 population size decreases with increasing dCas9 copy number via a mono-exponential decay, indicative of simple disassociation kinetics.